
Case Studies
At TME Scientific, we specialize in generating robust, genetically stable patient-derived xenograft (PDX) models that closely mirror primary tumor biology. Notably, our CEO, Dr. Hongwei Cheng, was a lead researcher on the landmark study by Chapuy et al., (2016) which demonstrated how DLBCL PDX models can drive functional drug discovery—a capability we now deliver directly to our clients.
PDX Model Generation & Expansion
Using freshly resected DLBCL biopsies, tumor samples were implanted under the renal capsule of NSG mice. From 28 attempts, 9 stable PDX lines were established and expanded across ≥5 passages—highlighting the feasibility of long-term in vivo modeling.
Model Characterization Included:
Clonality & EBV Status: EBV-negative, clonally confirmed via IgH PCR.
Phenotyping: IHC for markers such as CD20, CD10, MUM1, BCL6, BCL2, and MYC (Figure 2).
Transcriptomic Subtyping: RNA-seq classified tumors into ABC vs. GCB types and BCR vs. non-BCR subtypes.
Genomic Profiling: Whole-exome sequencing identified DLBCL-relevant mutations (e.g., MYD88, CD79B, EZH2) and hallmark translocations (e.g., IgH-MYC, IgH-BCL2).
Genetic Stability: Primary tumors and matched PDXs showed strong concordance in mutation profiles and allele frequencies.
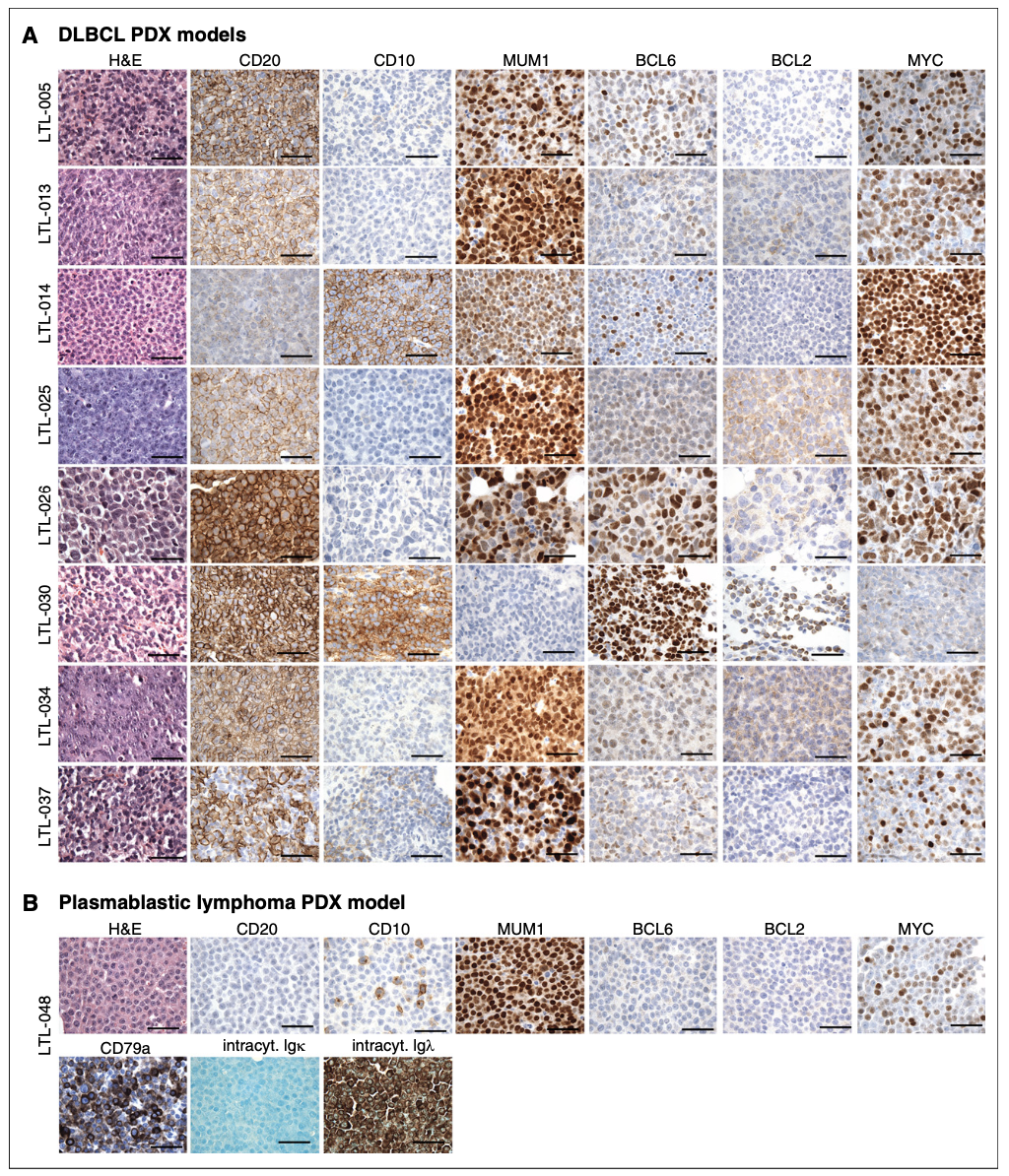
Figure 2. IHC characterization of all 9 PDX models. (A) IHC analyses of the indicated markers in all 8 DLBCL PDX models, which were consistent with the diagnosis of DLBCL. (B) IHC assessment of indicated markers in PDX model LTL-048, which is consistent with the diagnosis of PBL. Scale bars, 100 mm. See also Table 1. intracyt., intracytoplasmic.
Drug Response Correlation
Six of eight DLBCL PDXs were BCR-type and responded selectively to entospletinib, a SYK inhibitor. This demonstrated functional BCR signaling and validated the models' utility in evaluating targeted therapies (Figure 5).
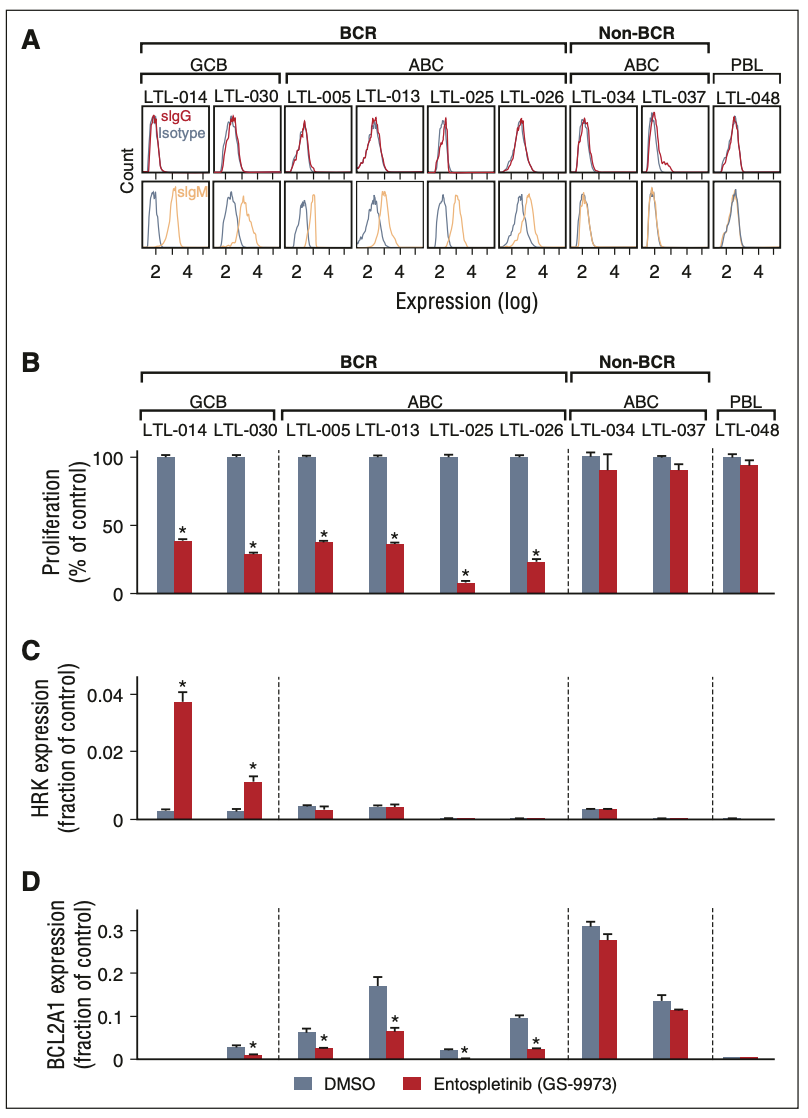
Figure 5. Analyses of cell surface immunoglobulin and BCR signaling in the LBCL PDX models. (A) Single-cell suspensions from each LBCL PDX model were gated for human CD45-positive cells and analyzed for surface immunoglobulin (IgG, red; IgM, orange; isotype, gray). CCC and COO subtypes of each model are indicated above the flow histograms. (B) Cellular proliferation of PDX tumor cell suspensions after chemical SYK inhibition with entospletinib (GS-9973) for 24 hours. (C,D) HRK (C) and BCL2A1 (D) transcript abundance following entospletinib (GS-9973) treatment. Experiments were performed in triplicate. A representative experiment of biological duplicates in shown. DMSO, dimethylsulfoxide. The P values were obtained using a Student t test (*P , .05).
Why It Matters
TME Scientific enables biotech and pharma partners to access PDX models that retain patient tumor fidelity, support mechanistic insights, and accelerate therapeutic validation—especially for lymphomas with defined genetic subtypes.

At TME Scientific, we support the integration of PDX models into co-clinical trial frameworks and personalized oncology. Our CEO, Dr. Hongwei Cheng, co-authored this foundational chapter (Cheng et al., 2017) detailing how PDX platforms can guide therapy decisions and model tumor evolution.
PDX Advantages & Implementation
PDX models are established by implanting patient tumors into immunodeficient mice. These models retain tumor heterogeneity, genetic stability, and therapy response, enabling longitudinal sampling and mechanistic studies of progression and resistance.
Co-Clinical Trials (Fig. 12.1): A dual approach using clinical trial patients and matched PDXs to optimize treatment selection. The figure illustrates the integrated workflow linking clinical sites, preclinical modeling, and bioinformatics for real-time translational research.
Therapy Resistance Modeling: PDXs simulate therapy-induced tumor evolution, such as neuroendocrine transformation in prostate cancer, providing insight into second-line treatment options.
Stable Genomic Profiles: Key mutations (e.g., KRAS, PIK3CA) are preserved across passages, ensuring reproducibility for drug testing.
Personalized Medicine Pipeline (Fig. 12.2): The timeline and workflow for PDX-guided treatment decisions are outlined—from tumor engraftment to drug efficacy testing in resistant models. PDXs help identify the most effective therapies for patients with relapsed or treatment-resistant disease.
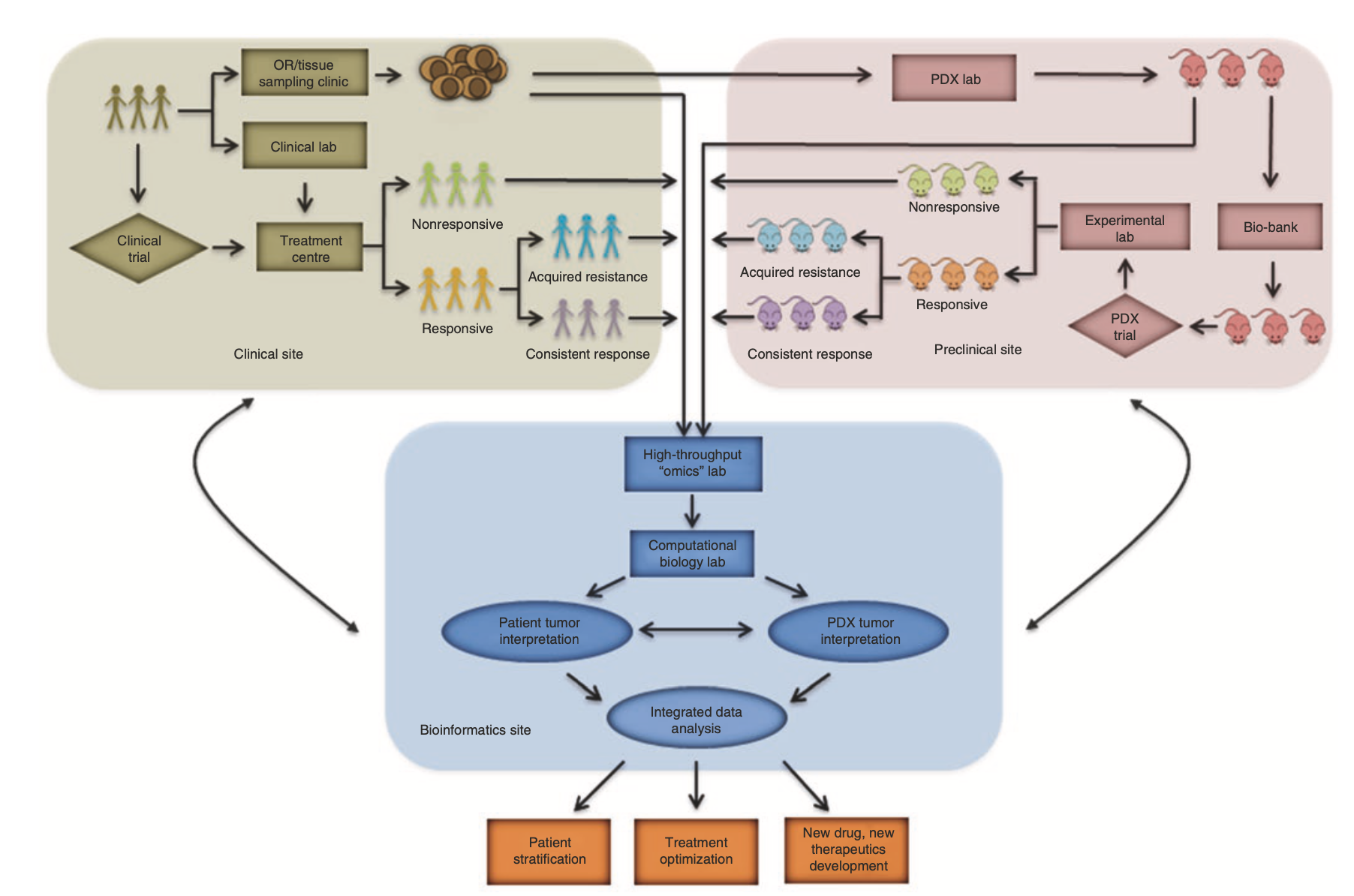
Fig. 12.2 Workflow and estimated timelines of PDX-assisted personalized medicine.
Why It Matters
PDX platforms offer a clinically relevant system for predictive drug testing, biomarker validation, and patient stratification, especially in oncology subtypes where genomic data alone may not be sufficient.
Fig. 12.1 Basic infrastructure and simplified workflow of a PDX-based co-clinical trial.

At TME Scientific, we apply advanced genomic tools to track tumor evolution in patient-derived xenografts (PDX). Dr. Hongwei Cheng, our CEO, co-authored this seminal Nature study (Eirew et al., 2015), which demonstrates how single-cell sequencing reveals the dynamic behavior of cancer clones post-engraftment—crucial for modeling drug response and resistance.
PDX Model Generation & Serial Passaging
Breast cancer tissues from 55 patients were serially engrafted into NSG and NRG mice, generating 30 xenograft lines with up to 16 passages over 3 years. Whole-genome sequencing and targeted deep sequencing were used to monitor somatic SNVs, structural variants, and copy number changes across timepoints.
Key Findings:
Clonal Selection on Engraftment:
All 15 patient samples exhibited clonal selection during initial engraftment. In many cases, minor clones in the primary tumor (<5%) became dominant in the xenografts.Longitudinal Clonal Dynamics:
Some xenografts underwent further clonal shifts over serial passages, particularly when initial selection was modest. These dynamics were highly predictable and reproducible in replicate transplants, indicating deterministic fitness advantages.Single-Cell Validation (Fig. 2):
For example, in case SA501, five distinct clonal genotypes were identified using single-cell sequencing and phylogenetic inference. This confirmed cascading subclonal evolution over passages X1 to X4, with genotype E eventually dominating the population.Clonal Replication in Replicates (Fig. 3):
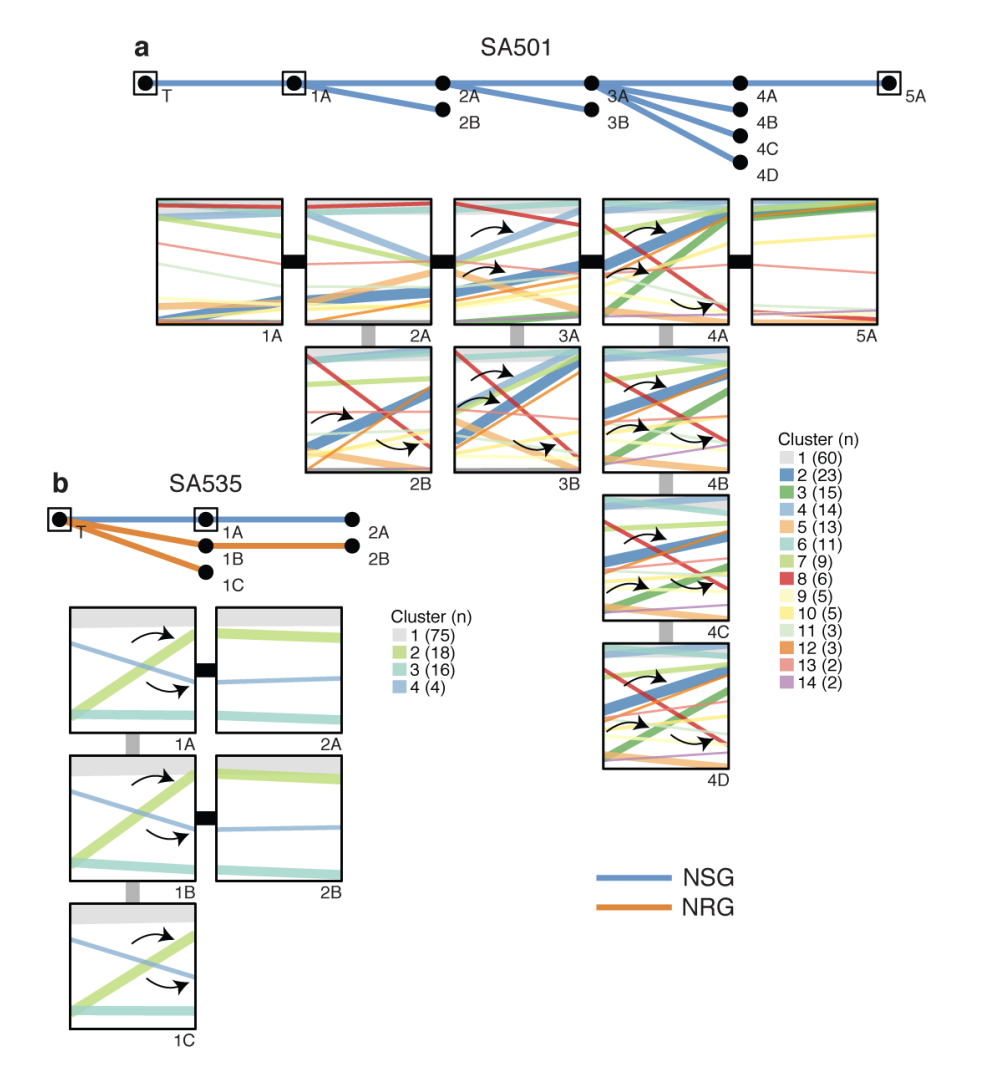
Clonal expansion patterns were consistently reproduced in independent transplant replicates (e.g., SA501, SA535), supporting the idea that genomic aberrations drive reproducible evolutionary trajectories in vivo.Predictive Genomics:
The study established that PyClone-inferred mutation clusters from bulk sequencing reliably match single-cell genotypes, offering a scalable framework for tracking tumor evolution.
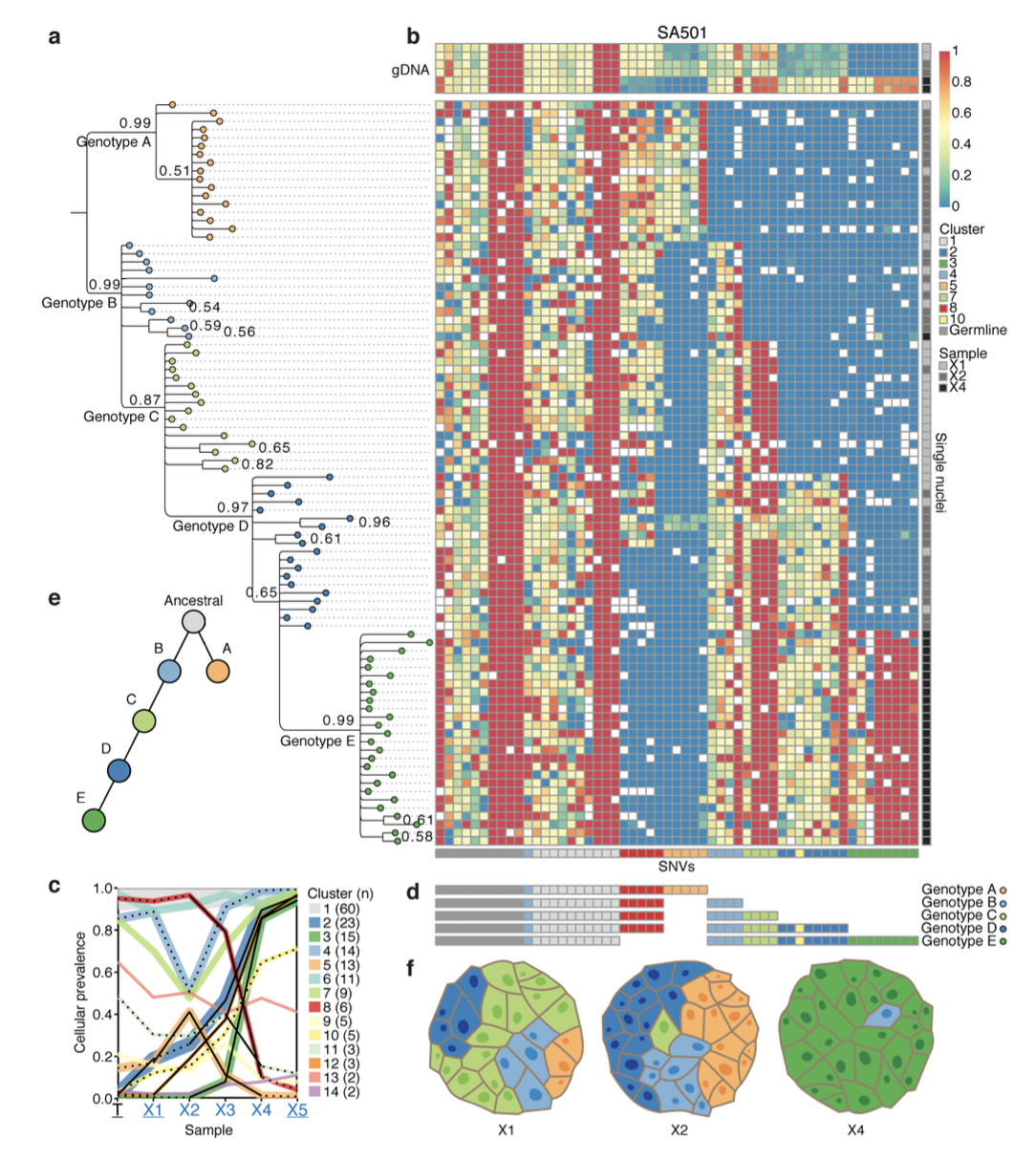
Figure 3. Clonal dynamics are reproduced in replicate transplants (1). a, b, Upper panels: Passaging history of SA501, SA535 showing transplants that resulted in successful xenografts. The host mouse strains (blue=NSG, orange=NRG) are indicated. Arrows show examples of parallel clonal dynamics of the same mutation cluster in multiple replicate transplants.
Figure 2. Single cell determination of clonal genotypes recapitulates population-based prediction of cascading subclonal evolution. a, Bayesian phylogenetic tree derived from multi-locus genotypes of individual nuclei, depicting cascading evolution. b, Heatmap depicting multilocus variant allele ratios (blue/ yellow/red corresponds to wild-type/heterozygous/homozygous loci). The cluster groupings (horizontal bar below horizontal axis) recapitulate the PyClone groupings inferred from bulk population measurements (c). d, Five consensus genotypes derived from high-probability splits in the phylogenetic tree. e, Schematic of the phylogeny derived from single cell genotyping depicts the sequential expansion of genomic subclones. f, Schematic representations of xenograft tumours X1, X2, and X4 based on single cell genotypes.
Why It Matters
These findings highlight the importance of monitoring clonal architecture in PDX studies to interpret therapy responses and resistance. At TME Scientific, we leverage this insight to support partners in oncology research, especially where genomic evolution shapes therapeutic outcomes.

References
Chapuy, B., Cheng, H., Watahiki, A., Ducar, M. D., Tan, Y., Chen, L., ... & Shipp, M. A. (2016). Diffuse large B-cell lymphoma patient-derived xenograft models capture the molecular and biological heterogeneity of the disease. Blood, The Journal of the American Society of Hematology, 127(18), 2203-2213.
Cheng, H., Liu, Z., Xue, H., Gout, P. W., & Shan, H. (2017). Application of PDX cancer models in co-clinical trials and personalized/precision medicine. Patient-derived xenograft models of human cancer, 177-192.
Eirew, P., Steif, A., Khattra, J., Ha, G., Yap, D., Farahani, H., ... & Aparicio, S. (2015). Dynamics of genomic clones in breast cancer patient xenografts at single-cell resolution. Nature, 518(7539), 422-426.
